Package managers
Managing software
1. Exploring an existing environment
Let’s open the Terminal app. Since Conda is not pre-installed in the Terminal app on UCloud, we will need to:
- Mount a drive with a pre-installed Miniconda setup.
- Run a bash script to add conda to the search path.
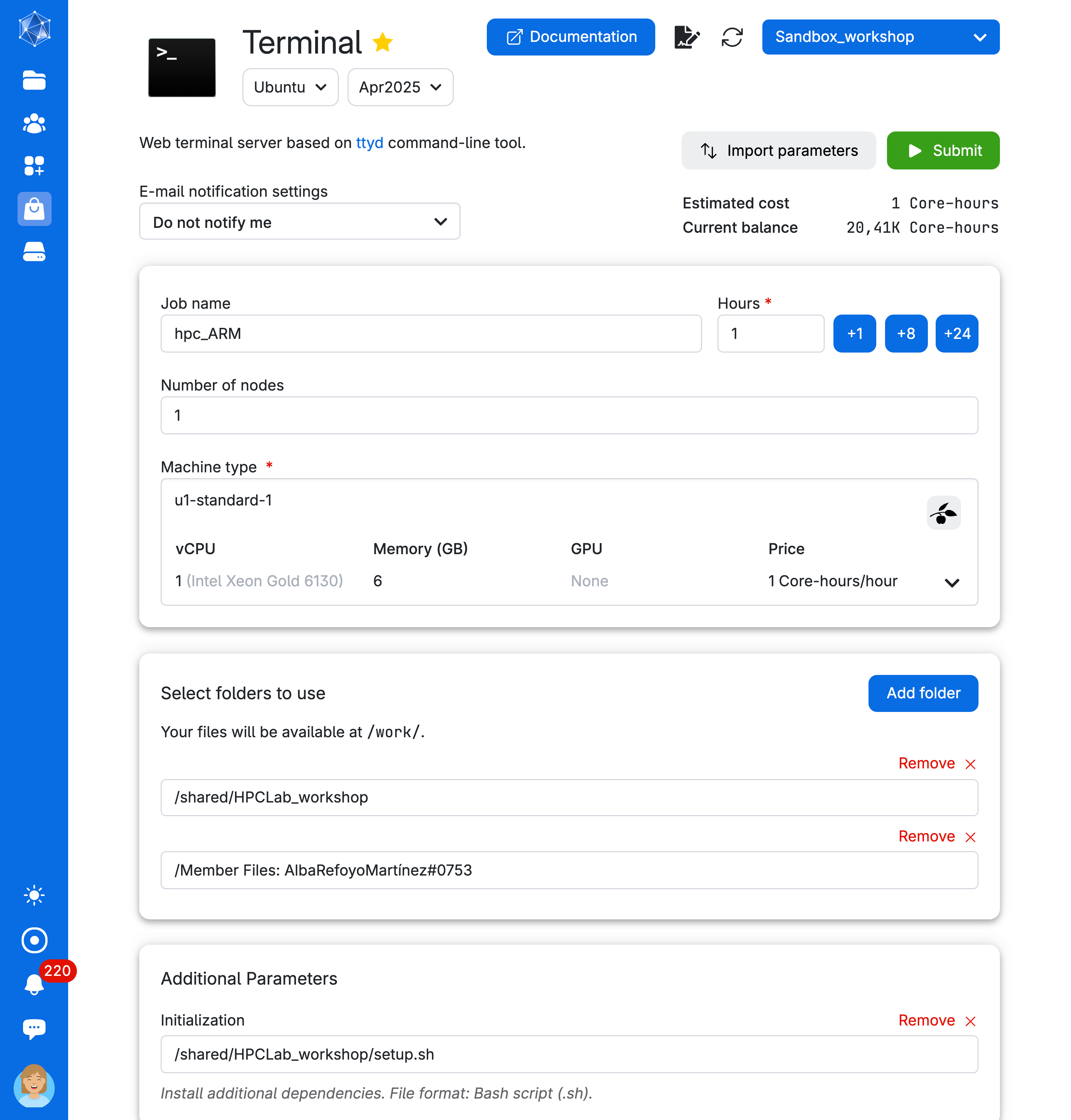
We will run the following commands to get familiar with Conda environments.
- What is the conda version?
conda info- Check the name of all the different environments available to you.
conda env list- Let’s explore one of the environments. To do this, let’s activate
hpclab-envenvironment
conda activate hpclab-env- How many packages are available in this environment?
conda list |grep -v '#' | wc -lImagine you need to share your environment with a collaborator so they can replicate your analysis. How do we do this?
- Export the environment specifications and save them to your personal drive (e.g.,
.yml)
conda env export --from-history > <yourname-hpclab>.yml- Deactivate the environment.
conda deactivate2: Building your conda environment
Let’s create a new directory in your personal drive:
cd <path-to-personal-drive>
mkdir envs Now, add a new environment using the argument --prefix PATH (e.g., /work/<YourNameSurname#xxxx>/envs/<name-env>). We need to do this as miniconda is installed in a directory that you don’t have write rights to.
Locally, you would typically run the command: conda create --name <myenv>
Always specify the location using --prefix PATH regardless of the UCloud app you use, especially if the app has conda pre-installed (e.g. Jupyterlab). The path must be on your personal drive or a shared drive with colleagues that you have access to. Otherwise, the environment won’t be saved, as you’re working within a temporary container instance.
- Create the environment
conda create --prefix /work/<NameSurname#xxxx>/envs/<test-env>- Confirm the environment location
Proceed ([y]/n)? y- Double-check that the new environment exists
conda env list- Activate the env
conda activate <myenv>- By default, we won’t have conda-forge or bioconda added to the environment. Let’s fix this.
conda config --remove channels defaults
conda config --add channels bioconda
conda config --add channels conda-forge- Add the latest Python version to the environment
Search available versions
conda search python Install the latest
conda install python=3.13.2 - Install samtools, a tool for manipulating DNA sequencing data
conda install -c bioconda samtools- Uninstall samtools
conda remove samtools - Deactivate your environment
conda deactivate